
Additionally, all nodes will be assigned the same position along the axis (the same radius), the same color, and node size.
This function will assign all nodes to a single axis. For example, if the list of interactions is given in the data frame denoted as “dataSet.ext,” we can create a hive object as: Given a network in an edge list format (data from column 1 and column 2 correspond to the interacting pairs of nodes), we can use the “ edge2HPD” command to create a hive object. For more information, see Network visualization – part 1 and Network visualization – part 2). I will also use the same node and edge properties: the degree of a node, betweenness centrality of a node, Dice similarity of two nodes, and the coappearance weight. To demonstrate how this works, I will use the same network I used to demonstrate network visualization in Cytoscape – the weighted network of characters’ coappearances in Victor Hugo’s novel “Les Miserables” ( LesMiserables.txt). As such, hive plots give users the ability to assess network structure using network properties they are interested in, as well as the ability to compare two networks based on the selected properties. Thus, using hive plots, users can create their own rules for a mapping between the network properties of interest and layout. Nodes are assigned to axes and position along the axis (denoted as the radius) based on their qualitative or quantitative properties, e.g., network structure, node, edge annotation, or any other meaningful properties of the network.
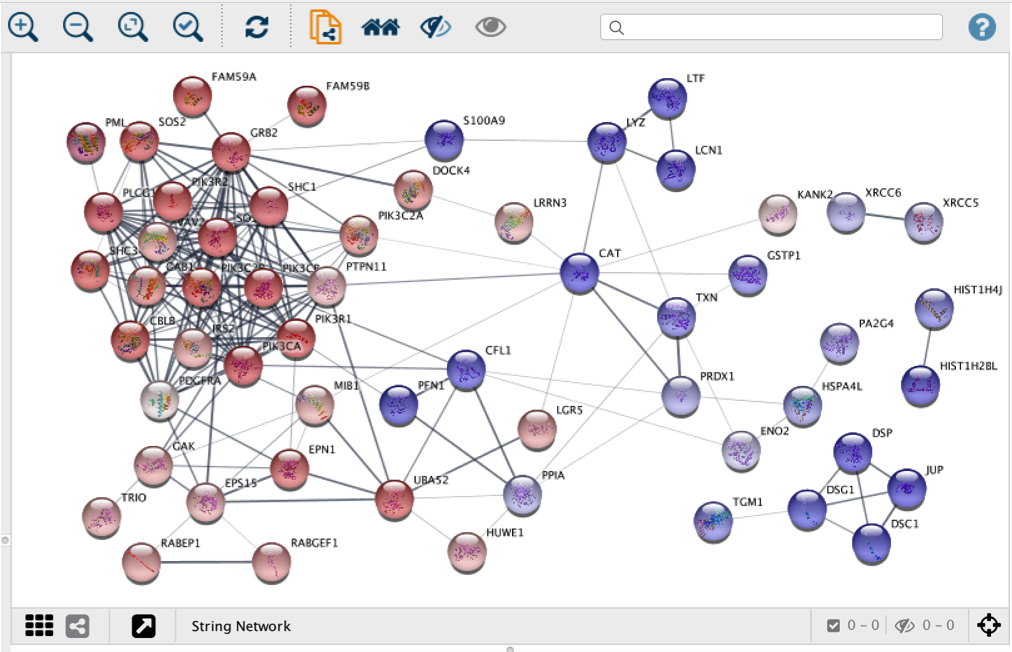
Hive plots map nodes onto radially distributed linear axes and edges between nodes are drawn as curved links that connect the axes. Thus, in the hive plots individual nodes and edges are not as important as individual elements, but as parts of a system. Additionally, standard network layouts generally work well for visualization of small/medium size networks, while visualization of large network often results in the “ hairball” network plots that lack identifiable structural patterns.Ĭonversely to standard network plots (i.e., layout algorithms), the goal of hive plots is to capture and expose both trends and patterns in network structure that arise from large number of nodes and edges, rather than solely representing network structure in the form of node-edge diagrams. For this reason, it is hard to assess how similar (or different) two networks are solely based on their resulting layouts/plots. Clearly, the resulting plots are sensitive to changes and even a small change in underlying topology can lead to a change in the final layout. The layout algorithms determine node (and edge) positions based on various criteria, e.g., the number of direct interacting partners, smallest number of edge crossings, or similar edge length between all nodes. The concept of hive plots is fundamentally different from the Cytoscape and Gephi plots.Ĭytoscape and Gephi use a number of layout algorithms to plot networks as node-edge diagrams in the Euclidean plane. In the third part of “how to quickly visualize networks directly from R” series, I’ll write about the hive plots and “ HiveR” package.
